Chorionic villus sampling (CVS) and amniocentesis are ways to collect a sample, not tests in themselves.
What matters is the type of genetic test performed on the sample: PCR, FISH, karyotype, microarrays, exome, whole genome sequencing (WGS), gene-specific sequencing


Diagnostic testing (CVS or amniocentesis) examines fetal genetic material directly and can provide a definitive diagnosis for the conditions tested, rather than a risk estimate. The choice between CVS (from 11 weeks) and amniocentesis (from 15 weeks) depends on timing and the specific test required.
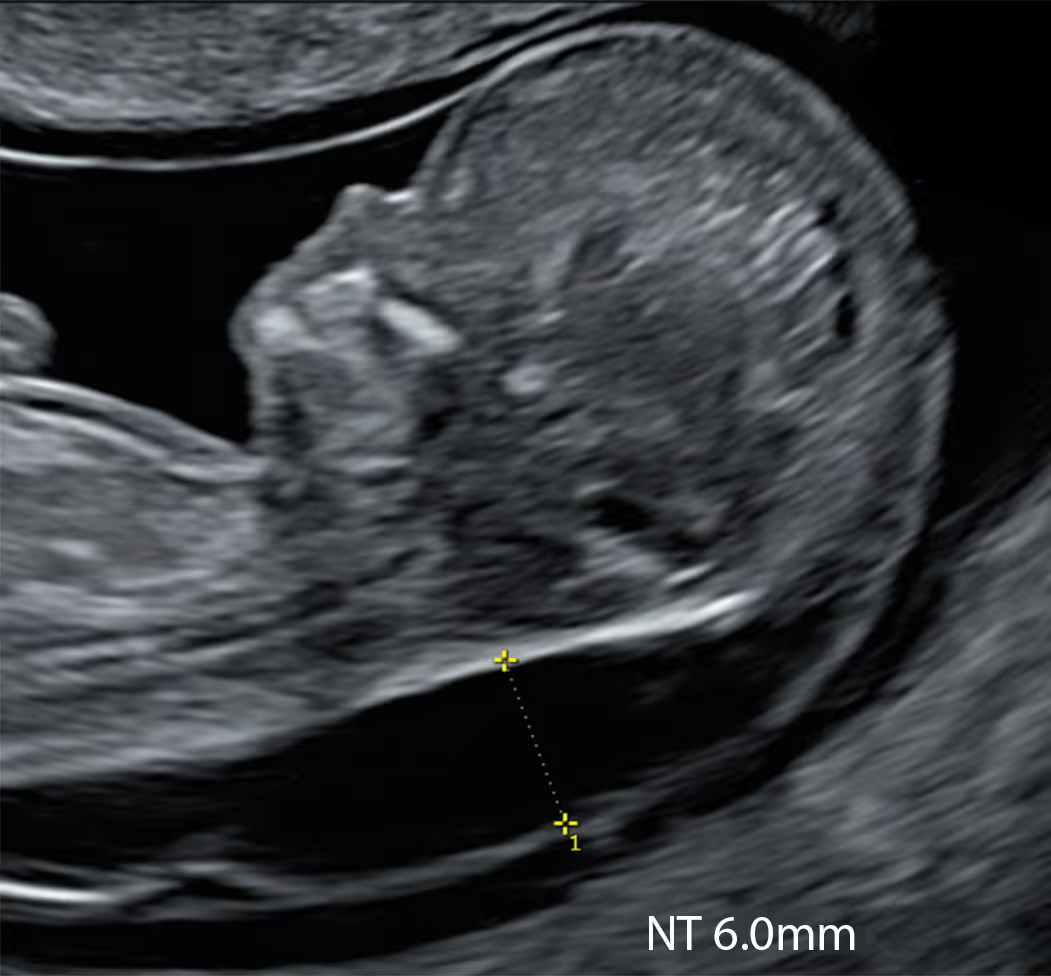
Increased nuchal translucency (NT) is a phenotype linked to a broad differential, many causes being rare, sporadic and de novo. To be truly diagnostic rather than merely invasive, the genetic assay must address the full differential, including monogenic disorders. A test that does not evaluate monogenic disease is not diagnostic.
Twenty years ago, karyotyping was considered diagnostic. By 2026, a truly diagnostic genomic work-up for high NT should include exome or whole-genome sequencing (WGS).
Genomics alone cannot exclude isolated structural anomalies; a targeted, expert ultrasound is still required to identify or rule these in or out.
Chorionic Villus Sampling (CVS): From 11 weeks, a tiny placental sample (chorionic villi) is taken for chromosome and gene tests. It is placental DNA, not fetal. Small miscarriage risk.
Amniocentesis: From 15 weeks, a small amount of amniotic fluid with fetal cells is taken for chromosome and gene tests. Small miscarriage risk (usually a little lower than CVS).
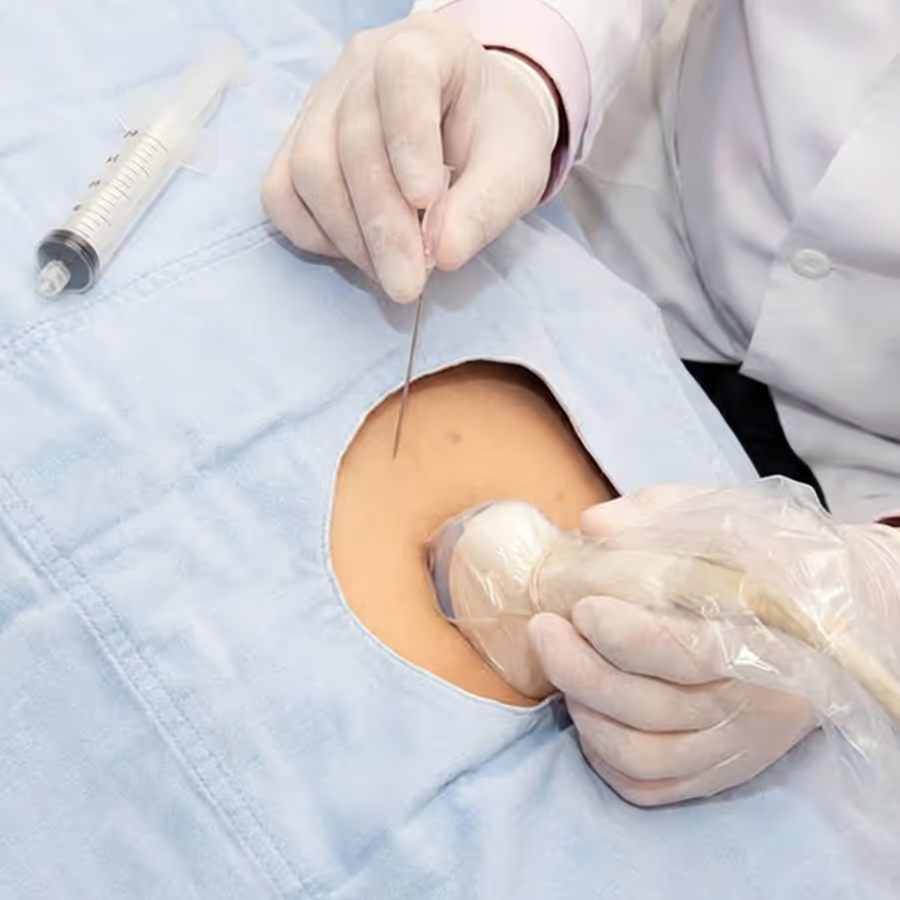
Most invasive prenatal tests are performed through the maternal abdomen (transabdominal) under continuous ultrasound guidance in a clinic setting.
After cleaning the skin, the clinician inserts a thin needle into maternal abdomen (tummy) to obtain a sample: CVS takes a tiny piece of placental tissue; amniocentesis draws a small amount of amniotic fluid. The needle is guided away from the baby and the placenta is approached only as needed; the whole procedure typically takes a few minutes.
Fetal heartbeat is checked before and after, and patients are advised to rest for the remainder of the day. Mild cramping or spotting can occur, and there is a some procedure-related risk of miscarriage, or infection.
Learn More about the SMART Protocol

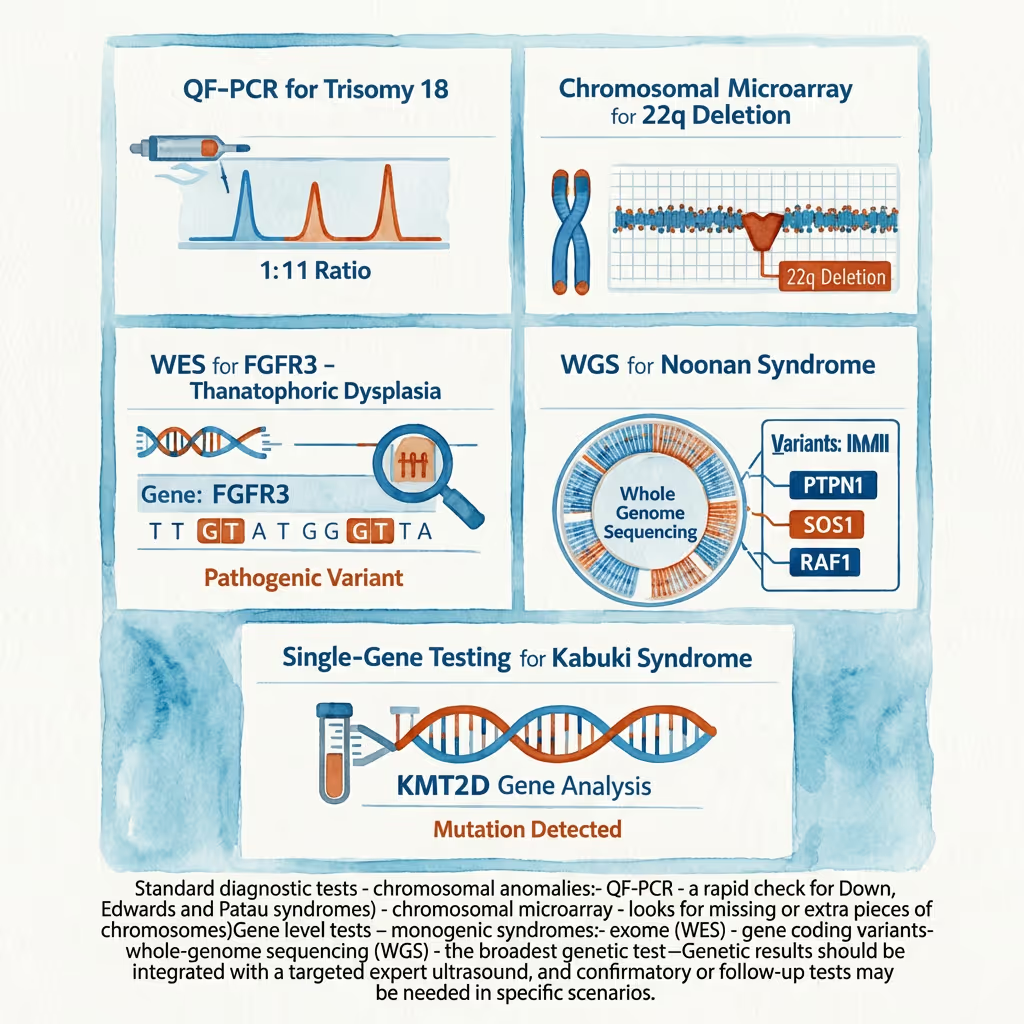
Standard diagnostic tests - chromosomal anomalies:
- QF-PCR - a rapid check for Down, Edwards and Patau syndromes)
- chromosomal microarray - looks for missing or extra pieces of chromosomes)
Gene level tests - monogenic syndromes:
- exome (WES) - gene coding variants
- whole-genome sequencing (WGS) - the broadest genetic test
Genetic results should be integrated with a targeted expert ultrasound, and confirmatory or follow-up tests may be needed in specific scenarios.

Learn More about the SMART Protocol

QF-PCR is a rapid assay performed on CVS or amniotic fluid for fetuses with increased NT. It provides a fast result (24-72 hours) for common aneuploidies: Down syndrome (T21), trisomies 18 and 13, and sex chromosome abnormalities, and can also suggest triploidy. Results can be affected by maternal cell contamination. In practice, QF-PCR is used as a rapid first-line test and is paired with microarray and, when indicated, exome or whole-genome sequencing for a truly diagnostic work-up.

Learn More about the SMART Protocol

Chromosomal microarray (CMA or aCGH/SNP array) is a standard genetic test that detects microdeletions and microduplications (missing or extra pieces of chromosomes). Microarray results usually return in about 1-2 weeks.
CMA does not detect most single-gene sequence variants (monogenic syndromes) and has other limitations.
In practice, it’s a first-line diagnostic test for increased NT when aneuploidy is excluded by QF-PCR or when a scan suggests a structural anomaly.

Learn More about the SMART Protocol

Whole-exome sequencing (WES) targets the protein-coding regions (exons) of ~19,000–20,000 human genes, which make up only ~1–2% of the genome yet harbour the great majority of known Mendelian disease-causing variants (often quoted ~85%).
Some services instead run a “clinical exome”, focusing on roughly 4,000–7,000 disease-associated genes, it is cheaper and quicker.
Because many causes of increased NT are monogenic and spread across thousands of genes, exome examines the coding genome to find diagnoses that QF-PCR/CMA miss and to guide prognosis and management.

Learn More about the SMART Protocol

Whole-genome sequencing (WGS) reads almost all of the fetal genome (≈3.1–3.2 billion bases), covering both coding and non-coding DNA regions, and often includes the mitochondrial genome.
It detects single-gene variants, small indels, and many copy-number and structural variants, offering broader coverage and more uniformity than exome. WGS still has limits, so a normal result does not exclude all disease.
WGS is the most expensive genetic test for increased NT. It offers a broader yield than exome testing, especially valuable in this situation, where the differential diagnosis of genetic syndromes is wide.

Learn More about the SMART Protocol

After a high-chance monogenic result on extended NIPT for increased NT, confirmation is done with targeted single-gene testing on CVS or amniotic fluid: PCR/Sanger sequencing to verify the specific SNV/indel flagged, or a focused NGS gene panel when several genes could explain the phenotype. This is diagnostic (not a screen), often run as a trio to assess inheritance, and provides definitive confirmation to guide prognosis and counselling.
QF-PCR (rapid aneuploidy screen): CVS/amnio; fast (same day–48 h) detection of T13, T18, T21 and sex-chromosome aneuploidy; can suggest triploidy.
Karyotype (G-banded): Whole-chromosome changes and large/balanced rearrangements; limited resolution; slow.
Chromosomal microarray (CMA; aCGH/SNP array): Genome-wide CNVs (~50–100 kb+); SNP arrays also assess ROH/UPD; cannot detect most single-gene variants or balanced rearrangements.
Targeted FISH: Rapid confirmation of suspected aneuploidies/rearrangements or specific microdeletions; locus-specific.
MLPA / MS-MLPA: Targeted exon-level CNVs; methylation/imprinting disorders (e.g., PWS/AS, BWS/SRS).
Single-gene testing (PCR/Sanger/NGS panels): For known familial variants or phenotype-driven monogenic disorders.
Exome sequencing (WES, ideally trio): Coding regions; SNVs/indels and some CNVs; phenotype-guided interpretation.
Whole-genome sequencing (WGS, ideally trio): Coding + non-coding; SNVs/indels/CNVs/selected SVs; best single comprehensive assay; may still miss repeats/methylation.
Low-pass (shallow) WGS: Genome-wide CNV/aneuploidy assessment; not suitable for monogenic variant detection.
Mitochondrial genome sequencing: mtDNA variants with heteroplasmy quantification; trio/segregation useful.
UPD studies (SNP array/STRs): Confirms uniparental disomy where suspected (imprinting/ROH patterns).
Quality checks: Maternal cell contamination and confined placental mosaicism assessment (STRs/array metrics); essential for result validity.
Both CVS and amniocentesis are considered very safe when performed by an experienced operator, and the difference in procedure-related miscarriage risk is small. In modern practice, the additional risk from either procedure is usually quoted as well under 1%, and often around or below about 0.5%, although exact figures vary between centres and depend on operator experience and individual pregnancy factors.
In real life, the bigger difference is timing. CVS is done earlier (typically around 11–14 weeks), so it can give an answer sooner. Amniocentesis is done later (from about 15 weeks), which can feel like a longer wait but has the advantage that it tests amniotic fluid rather than placental tissue.
A practical point is that CVS samples the placenta, and in a small minority of cases the placenta can have a genetic pattern that is not the same as the baby (placental mosaicism). If that happens, an amniocentesis may be recommended afterwards to clarify the result. So the “safest” choice is usually the one that fits your gestation and gives you the most reliable answer in your situation, performed by a highly experienced team.
Diagnostic genetic testing can help in three main ways: it can clarify the underlying cause, guide decisions and planning, and provide information for future pregnancies. When increased NT is accompanied by multiple severe anomalies, the chance of an underlying genetic problem is higher than with isolated increased NT. A diagnostic result may identify a chromosomal condition (such as a trisomy), a clinically significant microdeletion or microduplication, or a single-gene disorder that explains the pattern of findings. Even if the result is “normal”, that information is still useful because it narrows the range of likely causes and can help your team focus on the most appropriate follow-up, counselling and recurrence risk assessment.
Knowing the diagnosis can make counselling more accurate. It can help predict prognosis more reliably, avoid uncertainty, and sometimes prevent additional tests that are unlikely to add value. It can also guide pregnancy management, including the most appropriate scan schedule, whether specific organs need targeted assessment (for example detailed cardiac evaluation), and where the safest place is for ongoing care and delivery if continuing the pregnancy is being considered. If termination is being considered, a diagnosis can help you understand whether the condition is sporadic or inherited, and it can be important for closure and documentation.
Do you need the test? It is not mandatory, but it is usually strongly recommended in this situation because it often provides the most definitive explanation and the most useful information for both current and future care. If you choose to proceed, the most informative approach typically includes chromosomal microarray on CVS or amniocentesis, and in many cases additional gene-based testing such as a targeted panel or exome/genome sequencing, especially if microarray is normal or the anomaly pattern suggests a single-gene syndrome.
The decision ultimately depends on what information you want, how it might affect your choices, and your personal circumstances, and a discussion with fetal medicine and a genetic counsellor can help you choose the right level of testing.
It’s a very reasonable concern. QF-PCR and microarray are excellent for common trisomies and many chromosomal imbalances (including some microdeletions/duplications), but they do not rule out most monogenic syndromes that can sit behind an increased NT, such as RASopathies (Noonan spectrum), some skeletal dysplasias and other single-gene disorders. The key point is that you have already done the invasive part, so in many cases you can build on the same CVS sample rather than starting again.
The first step is to confirm exactly what material is still available from the CVS and what the laboratory has stored (DNA extract, cultured villi/cells, or remaining sample). Ask your team to ensure the sample (or extracted DNA) is retained, because that can be used for additional testing. Then arrange a discussion with a consultant clinical geneticist, because the most appropriate next genetic test depends on the ultrasound picture (isolated increased NT versus additional findings) and the family history.
From there, you typically have three realistic “next test” directions. One is a targeted gene panel, most commonly a RASopathy panel if the pattern fits, which can be quicker and more focused. The second is whole-exome sequencing, which looks broadly across many genes and is often chosen when the ultrasound findings are worrying or not clearly pointing to one syndrome. The third is whole-genome sequencing, which is the broadest option and can sometimes pick up variants that exome may miss, depending on the laboratory’s pipeline and reporting strategy. In many situations, testing works best as a trio (baby plus both parents), because it improves interpretation and speed of decision-making.
In the UK, there is a private pathway where a consultant clinical geneticist can request that existing CVS DNA (or remaining material) is sent to a private laboratory for exome or whole-genome sequencing, particularly when there is an abnormal ultrasound finding such as increased NT. If the CVS material is insufficient or the result raises a question about placental-only findings, your team may sometimes discuss whether an amniocentesis later would be more informative for follow-on testing, but often the CVS sample is adequate for monogenic testing if enough DNA is available.
If you want a non-invasive “bridge” while arranging definitive testing, an extended NIPT that includes a monogenic panel can sometimes be considered, and KNOVA NIPT can be particularly relevant here, especially its monogenic-only option. The main value in your situation is not to replace diagnostic testing, but to potentially flag a specific syndrome that can then be directly confirmed on the CVS DNA you already have, without another invasive procedure. That said, if you can access exome or genome on the CVS sample, that remains the more definitive route.
Most of the time, CVS or amniocentesis can be performed, but not always on the first attempt and not always on the same day. Whether it can be done depends on gestational age, the position of the placenta and baby, your anatomy, and safety factors on the day. Sometimes the safest option is to postpone for a few days, choose a different route (for example transcervical versus transabdominal CVS), or switch from CVS to amniocentesis later.
There are situations where an invasive test may be temporarily unsuitable. Examples include significant vaginal bleeding, signs of infection or fever, certain technical difficulties such as an inaccessible placenta, a uterus that is difficult to access because of fibroids or scarring, or if the pregnancy is too early for safe sampling. In multiple pregnancies the procedure is still possible, but it is technically more complex and needs specialist expertise; in some twin situations amniocentesis may be preferred.
Occasionally, even with an experienced operator, it may not be possible to obtain a good sample on the day, or the laboratory may report that the sample was insufficient or did not grow well (for CVS culture), and a repeat attempt or an alternative approach may be needed. The safest principle is that the test should only be performed if the operator can access the target safely under clear ultrasound guidance; if not, delaying or choosing a different method is the right decision.
A retroverted uterus is common, and in early pregnancy it can make CVS technically harder because the angle and access to the placenta are less favourable. In many cases, the situation improves simply with time as the uterus rises out of the pelvis and the placenta becomes easier to reach, so postponing is often the safest plan rather than forcing a difficult procedure.
There are a few practical things that can sometimes help, but they are not guaranteed. A repeat attempt a few days later or at a slightly later gestation is often enough. Some operators may try a different approach, such as a transabdominal rather than transcervical route, or vice versa, depending on where the placenta sits. Occasionally, having a comfortably full bladder can improve the ultrasound window and change the angle of access, but you should only do this if your clinic advises it, because too full can make things worse.
While waiting, doing NIPT can be a sensible “bridge”, especially if you want some information quickly and safely. KNOVA can be particularly useful in an increased NT scenario because, beyond the common trisomies, it can also screen for a broader range of genetic conditions linked to increased NT, including a panel of selected monogenic syndromes; in some pathways it can also be done as a monogenic-only panel if that is the main concern. If KNOVA suggests a specific condition, you can then plan targeted confirmatory testing once CVS becomes possible, without losing time.
The key is not to let the pathway stall: book the repeat specialist scan and the next available CVS slot, do NIPT promptly if you want interim risk information, and have a clear plan for what you will do with each possible result. If CVS remains technically difficult, amniocentesis a little later is usually an alternative route to definitive testing.
Yes, you can choose to do NIPT now and delay an invasive test, and many people do this because NIPT is safe and can quickly reduce uncertainty. The key point is that when the NT is increased, this is not a “routine screening” situation, so a basic trisomy-only NIPT is often too limited. Increased NT is linked not only to Down/Edwards/Patau syndromes, but also to a wider range of chromosomal changes, selected microdeletions and, importantly, single-gene (monogenic) syndromes such as RASopathies, Kabuki syndrome and some skeletal dysplasias.
This is exactly where an extended NIPT such as KNOVA is most relevant. Compared with basic NIPT, KNOVA can screen more broadly for genetic conditions that are known to sit behind increased NT, including a panel of selected monogenic syndromes, and this can meaningfully reduce uncertainty while you continue expert ultrasound follow-up and consider whether you want definitive testing later. It still cannot cover every possible condition and it remains a screening test, but it is better aligned with the types of genetic diagnoses that become more important when NT is increased.
Waiting until 20 weeks has trade-offs. You may learn more from ultrasound over time, including the detailed anatomy scan and often a better assessment of the heart, which can clarify prognosis. The downside is that if you decide later that you need a definitive genetic answer, amniocentesis at that stage can push final results later, which can make decision-making harder and more time-pressured.
A practical approach many families choose is: do KNOVA now (rather than basic NIPT), continue staged expert scans, and keep the option of invasive diagnostic testing open if KNOVA is high chance, the NT or nuchal fold (NF) remains significantly raised, or new anomalies appear.
It depends on what testing has been done (or is planned) on your CVS sample. If the CVS is being analysed with a comprehensive genetic approach such as chromosomal microarray plus whole-exome sequencing (WES) or whole-genome sequencing (WGS), then NIPT usually adds nothing, because you are already getting more definitive, diagnostic information directly from fetal/placental tissue. However, in many cases CVS testing is limited to rapid aneuploidy testing (for example QF-PCR) and/or microarray, which mainly assess chromosome problems and may not look for single-gene (monogenic) syndromes unless specifically requested. This matters because increased NT is strongly associated with a range of monogenic conditions, particularly RASopathies (Noonan spectrum), some skeletal dysplasias and other severe syndromes. In that scenario, KNOVA can be relevant, especially the option that focuses on its monogenic panel, because it can screen for a defined set of single-gene disorders that may not otherwise be assessed.
A practical advantage is that if KNOVA flags a specific monogenic condition, you can often use the DNA already obtained from the CVS to confirm (or refute) that exact finding, without needing another invasive procedure. The key is to review exactly what is being tested on your CVS sample and decide whether adding targeted monogenic screening (or arranging diagnostic gene testing on the CVS DNA) is the most efficient next step.
For medical professionals who want to learn more about the SMART NT approach, access training, and join our clinical network.