Extended NIPT goes beyond trisomies 21/18/13 to include additional chromosomes, key microdeletions, and selected single-gene syndromes - especially useful in increased NT, where many causes lie beyond the basic panel.

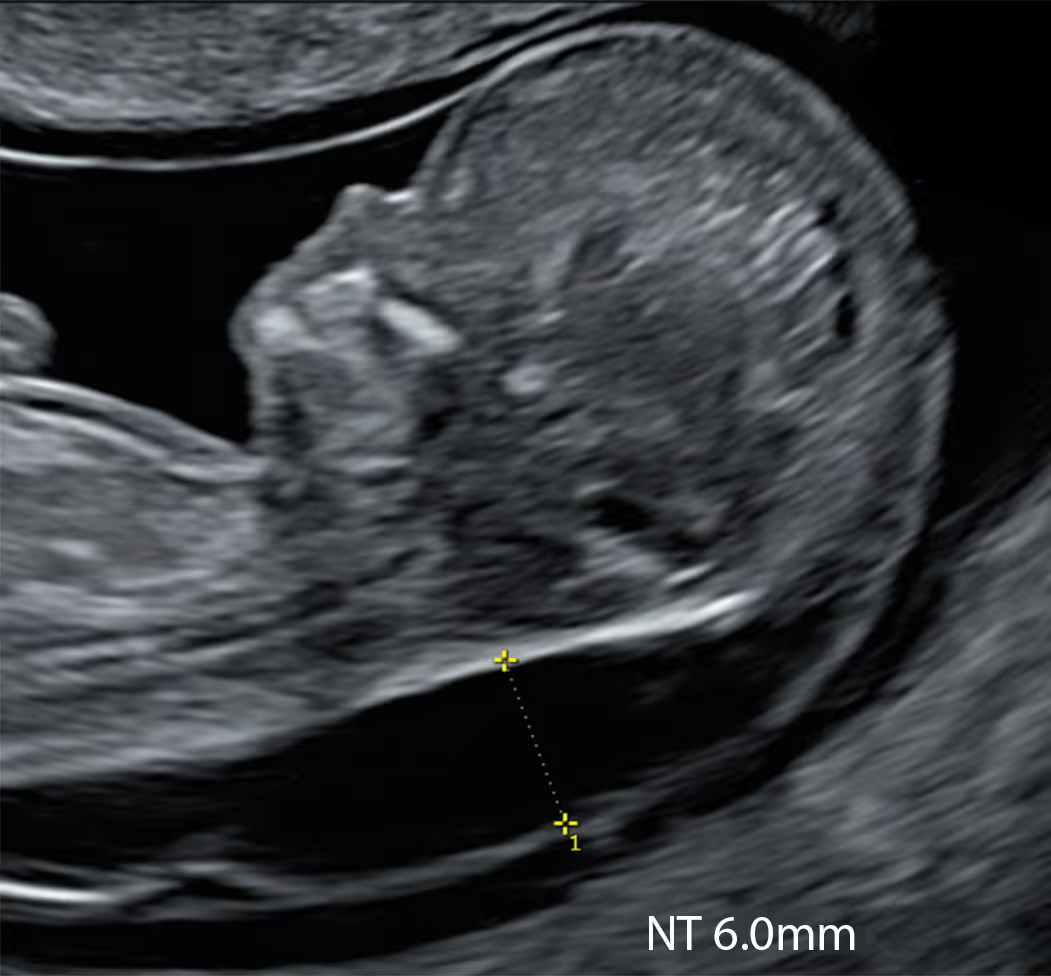

Increased NT (iNT) is strongly associated with de novo chromosomal and monogenic conditions, so the best NIPT for iNT must cover both - aneuploidies/CNVs and key single-gene syndromes. Many so-called extended panels add little beyond the basic trisomies; in iNT their yield is often similar to basic NIPT.
Much of the confusion comes from marketing labels (“complete”, “plus”, “premium”, “extended”) and the misused term “genome-wide (GW) NIPT”, which does not read the genome or test individual genes - it samples cfDNA at shallow depth to flag large chromosomal imbalances.
Because most “extended” NIPTs add only sex-chromosome aneuploidies and a few microdeletions, their utility in iNT is limited: the only consistently iNT-associated conditions they capture are monosomy X (45,X) and 22q11.2 deletion.
We have reviewed current options and identified a small subset of genuinely fit-for-purpose NIPT panels for the high-NT scenario. KNOVA by Fulgent (US) is the best option.
Even the most advanced NIPTs, are screening tests - not diagnostic. Any high-chance result should be confirmed with an invasive diagnostic test. Genetic testing from CVS/amniocentesis should be tailored to the NIPT result (e.g. karyotype/CMA or gene-specific sequencing).
Despite the many basic NIPTs with microdeletion add-ons, only four tests are truly fit for the high-NT setting. Among them KNOVA by Fulgent stands apart for genuine, sophisticated technology and class-leading coverage of increased NT-associated conditions.
KNOVA offers the broadest coverage across all other extended NIPT options. It screens for 6 autosomal trisomies, 4 sex‑chromosome aneuploidies, 12 microdeletions and 56 single genes associated with multiple monogenic conditions. A significant proportion of the conditions targeted by KNOVA are linked to increased NT (iNT), making it our NIPT of choice in this setting. It also offers broader coverage, a fast turnaround (~2 weeks), and a very low no-call rate (<1%).

Learn More about the SMART Protocol

PrenatalSafe Complete Plus can also be used in the setting of increased NT. It runs two parallel pipelines: a genome-wide chromosomal cfDNA screen (all chromosomal aneuploidies, large deletions/duplications, and nine targeted microdeletions) and a separate monogenic panel (25 single-gene disorders) processed in another division of the laboratory. While less integrated and, in our view, less comprehensive than KNOVA, it remains useful; particularly in vanishing-twin and donor-egg pregnancies.

Learn More about the SMART Protocol

Panorama with its microdeletion add-on offers little advantage over basic NIPT; however, pairing Panorama with Vistara (monogenic NIPT) can be useful in the iNT setting. The combination is relatively expensive and, even together, covers far fewer conditions than KNOVA: four chromosomal aneuploidies, sex-chromosome anomalies, five microdeletions, and around 30 single-genes associated with various monogenic disorders.

Learn More about the SMART Protocol

NIFTY Mono (BGI) may offer a more expansive platform; however, due to ongoing, well-publicised concerns (data-governance/national-security issues in the US/UK), we will not describe or endorse it further until the situation is resolved.
“Extended” or “advanced” NIPT are marketing-style umbrella terms and, on their own, they don’t tell you what is actually being tested.
When NT is increased, the words “extended” or “advanced” NIPT can be particularly misleading, because in this scenario the key question is not “more or less advanced”, but “does this test cover the types of conditions that increased NT is actually linked with?”
In this situation it matters, because increased NT is linked not only to Down, Edwards and Patau syndromes, but also to a wider range of chromosomal changes (including some microdeletions/microduplications) and single-gene (monogenic) syndromes such as RASopathies (Noonan spectrum) and some skeletal dysplasias. A trisomy-only NIPT is excellent for trisomy 21/18/13, but it does not address much of the remaining genetic risk that comes with increased NT.
Rule of thumb for patients: If your baby has increased NT, don’t ask “is this an advanced NIPT?” — ask “exactly which conditions does this NIPT screen for, and does it include a single-gene (monogenic) panel and microdeletions?”
Monogenic screening is different from chromosome screening, and this is especially relevant when NT is increased. These tests are designed to detect specific pathogenic DNA variants linked to serious single-gene syndromes, often aiming to pick up a new (de novo) fetal mutation from placental DNA fragments circulating in the mother’s blood. The underlying sequencing technology is highly advanced and similar to methods used in diagnostic genetics laboratories, so for many targets the analytical specificity is very high and true “technical” false positives are thought to be uncommon. In simple terms, if a pathogenic variant is reported, that DNA fragment was present in the blood sample.
The more common real-world complexity is not a laboratory “mistake”, but biology and interpretation. Because the mother’s DNA is always present in the sample, the test may sometimes detect a pathogenic or likely pathogenic variant in the mother (even if she is healthy or has only mild features). In that situation, the result can look like a “positive fetal screen”, but what it may actually mean is an inherited-risk scenario, where the baby could have up to a 50% chance of inheriting the variant depending on the gene and inheritance pattern. In some situations, paternal inheritance can also be relevant, but maternal DNA findings are the most frequent source of unexpected complexity.
For these reasons, any monogenic “high chance” result must be handled with careful counselling and confirmatory testing before it is treated as a diagnosis. It is also extremely important to arrange an expert targeted ultrasound to look for phenotype–genotype correlation, meaning specific ultrasound features that fit (or do not fit) the suspected condition. This can be complex, particularly in the increased NT setting, and it is best managed by a multidisciplinary team that includes fetal medicine specialists, genetic counsellors/consultant geneticists, and sometimes other relevant paediatric specialists.
That is why, before choosing tests such as KNOVA or Vistara, it is sensible to check that your provider has a clear pathway for counselling, rapid follow-up, and diagnostic confirmation if needed.
Both approaches are targeted rather than “screen everything”, which helps reduce unnecessary uncertainty and limits false-positive results compared with genome-wide screening. KNOVA is also based on very advanced genomic technology and, in practice, it can be faster and more cost-effective because it delivers broader coverage in a single pathway and a single report, rather than using a tandem approach.
In terms of scope, KNOVA includes 10 clinically relevant chromosomal aneuploidies, 12 microdeletions, and a single-gene panel of 56 genes associated with multiple conditions. By comparison, the Natera tandem approach (Panorama plus Vistara) typically covers 8 clinically relevant aneuploidies, 5 microdeletions, and 30 single genes. Another practical advantage of KNOVA is that you receive one integrated report rather than two separate reports from two different tests.
KNOVA is performed by Fulgent, a laboratory with a strong reputation and deep expertise in genetic testing, particularly in the field of rare diseases. This matters in the increased NT setting because many clinically important scenarios involve rarer genetic conditions where laboratory experience and interpretation pathways are crucial.
Overall, if KNOVA is available, it is the stronger and more efficient option, particularly when NT is increased and you want targeted coverage of conditions that overlap with increased NT. If KNOVA is not available in your country, Natera’s tests remain a reasonable alternative, as long as counselling is clear about what is covered, what is not, and how any high-chance finding would be confirmed.
In most increased-NT scenarios, the single-gene (monogenic) component is often the more useful “extra” compared with microdeletions. The reason is that after the common trisomies are considered, a significant proportion of the remaining genetic diagnoses linked to increased NT are single-gene syndromes, particularly RASopathies (Noonan spectrum) and certain skeletal dysplasias and other severe syndromes. These conditions are exactly the type of diagnoses that a monogenic panel is designed to screen for, and they can also fit well with an ultrasound-based pathway because there are often recognisable patterns that can be looked for on expert scanning (phenotype–genotype correlation).
By contrast, microdeletions are a mixed group in this context. A few specific microdeletions are relevant to increased NT (for example 22q11 deletion in the right clinical setting), but many clinically significant microdeletions and microduplications associated with increased NT are sporadic and not limited to a small “named syndrome” list. Extended NIPT microdeletion panels are usually restricted to selected syndromes and do not provide comprehensive genome-wide CNV detection. In addition, microdeletion screening tends to have a higher false-positive burden than trisomy screening, which can generate anxiety and lead to invasive confirmation anyway.
So, if you are choosing an extended NIPT specifically because NT is increased, the single-gene panel is often the part that adds the most clinically relevant extra coverage beyond trisomies, especially for Noonan-spectrum conditions and selected skeletal dysplasias. Microdeletion screening can still add value in some pathways, but it is best thought of as targeted and limited, and it does not replace diagnostic testing with microarray when you truly need comprehensive CNV assessment.
In general, the highest false-positive burden in extended NIPT comes from sex chromosome conditions, microdeletions, rare autosomal trisomies, and genome-wide/segmental chromosome imbalances. So if you are considering genome-wide (GW) “karyotypic NIPT” screening of all chromosomes, you can sometimes open a Pandora’s box.
There are two main reasons false positives are more common in these categories. First, these findings are much rarer than trisomy 21/18/13, so even a small amount of biological “noise” can lead to more false alarms. Second, NIPT analyses placental DNA, and the placenta can carry mosaic abnormalities that are not present in the baby. Maternal factors can also contribute, such as maternal mosaicism or benign maternal copy-number variants, and a vanishing twin can complicate interpretation.
You can’t completely avoid false positives in extended panels. Even if each individual condition has a small false-positive rate, when you screen for many conditions the overall chance of at least one false-positive result inevitably increases.
However, you can reduce the impact by choosing a well-curated, clinically focused panel rather than a “screen everything” approach. Avoid broad genome-wide, all-chromosome “karyotypic” NIPT unless there is a clear reason, because this tends to generate more uncertain or false-positive findings. Instead, prioritise panels that target well-validated chromosomal conditions and a carefully selected list of microdeletions and genes/syndromes with strong prenatal relevance and meaningful clinical actionability, ideally those associated with serious outcomes and/or well-recognised ultrasound patterns.
A good example of this approach is KNOVA by Fulgent, which is designed as a targeted, clinically curated extended panel rather than an indiscriminate “all chromosomes” screen. It combines screening for the common trisomies with selected additional categories (such as specific microdeletions and a defined single-gene panel), which is particularly relevant in the increased NT setting.
You can further reduce harm from false positives by combining extended NIPT with expert ultrasound (to look for phenotype–genotype correlation), ensuring proper pre-test counselling, and having a clear plan for confirmatory diagnostic testing (CVS/amnio) before any major decisions are made.
No. An increased NT is associated with a very wide range of genetic causes, including many chromosomal abnormalities, microdeletions/microduplications, and single-gene syndromes. In total, the spectrum includes around a hundred recognised genetic conditions and hundreds of possible underlying genetic variants, so no screening panel can cover everything.
Extended NIPT tests such as KNOVA can cover a much broader and more relevant set of conditions than basic trisomy-only NIPT, and in practice they can screen for many of the more common and clinically important genetic causes linked to increased NT. This can be particularly helpful because it goes beyond trisomies 21/18/13 and includes selected microdeletions and a panel of single-gene conditions that overlap with increased NT, such as RASopathies and some skeletal dysplasias.
However, even the best extended NIPT does not screen for all possibilities. In particular, it does not provide a comprehensive screen for all sporadic microdeletions and microduplications across the genome, which are a meaningful cause of increased NT and are better assessed with diagnostic testing and chromosomal microarray. It also does not cover the majority of autosomal recessive conditions and many X-linked disorders that can be associated with increased NT, because those conditions often require different testing strategies and, in many cases, parental carrier status information.
That is why, when NT is increased, extended NIPT can be a very useful part of the pathway, but it does not replace expert ultrasound follow-up and, when indicated, diagnostic testing such as CVS or amniocentesis with microarray and, in selected cases, exome or whole-genome sequencing.
There is significant misuse of the term “genome-wide (GW) NIPT” in papers, marketing and clinic websites. GW NIPT does not read the whole genome or analyse individual genes. It performs shallow, low-coverage counting of DNA fragments across chromosomes to look for large gains or losses of chromosomal material. Think of it as a broad map scan, not a line-by-line read of the genetic book.
What GW NIPT can screen: the common trisomies (21/18/13), often sex-chromosome changes, and in some labs selected rare autosomal trisomies or large segmental deletions/duplications.
What it cannot do: detect single-gene (monogenic) conditions or most small microdeletions/duplications, nor provide true whole-genome or diagnostic analysis.
Where single-gene (monogenic) NIPT fits: this is a separate, targeted cfDNA approach that screens specific genes/conditions (e.g., some recessive disorders or defined syndromes). Panels and performance vary by lab; many require known parental variants or are limited to a set list of genes. They are still screening tests - any high-chance result needs CVS/amnio for confirmation.
Take-home: call GW NIPT what it is - a chromosome-level cfDNA screen at shallow depth. Use it alongside ultrasound, and don’t confuse it with single-gene NIPT or whole-genome sequencing by diagnostic invasive test.
For medical professionals who want to learn more about the SMART NT approach, access training, and join our clinical network.